|
Rainbow Transgenic Flies
egfp-grna expressing vectors  Egfp Grna Expressing Vectors, supplied by Rainbow Transgenic Flies, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/egfp-grna expressing vectors/product/Rainbow Transgenic Flies Average 90 stars, based on 1 article reviews
egfp-grna expressing vectors - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
VectorBuilder GmbH
a grna expression vector targeting human mmp3 (cat. no. vb170623-1031qnn)  A Grna Expression Vector Targeting Human Mmp3 (Cat. No. Vb170623 1031qnn), supplied by VectorBuilder GmbH, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/a grna expression vector targeting human mmp3 (cat. no. vb170623-1031qnn)/product/VectorBuilder GmbH Average 90 stars, based on 1 article reviews
a grna expression vector targeting human mmp3 (cat. no. vb170623-1031qnn) - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Nature Biotechnology
grna-expression vectors  Grna Expression Vectors, supplied by Nature Biotechnology, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/grna-expression vectors/product/Nature Biotechnology Average 90 stars, based on 1 article reviews
grna-expression vectors - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Oligos Etc
grna expression vectors  Grna Expression Vectors, supplied by Oligos Etc, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/grna expression vectors/product/Oligos Etc Average 90 stars, based on 1 article reviews
grna expression vectors - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Amaxa
grna expression vectors  Grna Expression Vectors, supplied by Amaxa, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/grna expression vectors/product/Amaxa Average 90 stars, based on 1 article reviews
grna expression vectors - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
VectorBuilder GmbH
grna expression vectors for pgr targeting pgr640a  Grna Expression Vectors For Pgr Targeting Pgr640a, supplied by VectorBuilder GmbH, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/grna expression vectors for pgr targeting pgr640a/product/VectorBuilder GmbH Average 90 stars, based on 1 article reviews
grna expression vectors for pgr targeting pgr640a - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
VectorBuilder GmbH
grna expression vectors  Grna Expression Vectors, supplied by VectorBuilder GmbH, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/grna expression vectors/product/VectorBuilder GmbH Average 90 stars, based on 1 article reviews
grna expression vectors - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: Fly
Article Title: A simplified and efficient germline-specific CRISPR/Cas9 system for Drosophila genomic engineering
doi: 10.4161/fly.26828
Figure Lengend Snippet: Figure 1. Disrupting EGFP and mRFP genes using a germline-specific CRISPR/Cas9 system. ( A ) Scheme for generating Vasa-Cas9 transgenic flies and designing EGFP and mRFP gRNAs. The germline-specific Cas9 expressing vector, pVasa-Cas9, was integrated into the ZH-2A site on chromosome X mediated by the phiC31 integrase/attP/attB integration system. The resulting Vasa-Cas9 strain contains the germline-specific Vasa-driven Cas9 sequence and the eye promoter 3xP3-driven EGFP and mRFP sequences. Two gRNAs were designed to target mRFP and EGFP, respectively. Blue letters indicate the gRNA spacer sequences. The 3′ common regions of the gRNAs are represented with identical hairpin structures. Bold magenta labels the PAM motifs of the DNA substrates. Cyan triangles indicate the Cas9 cut sites. Key chromogenic residues are labeled red and green respectively. ( B ) EGFP and mRFP expression in G1 adult eyes. Representative images of G1 adult eyes from Vasa-Cas9 G0 flies injected with EGFP (top panels) or mRFP (bottom panels) gRNA-expressing vectors. Six flies are shown in each condition: those that retain fluorescent signals in top rows, and those showing a loss of fluorescent signals in bottom rows. EGFP signals are shown on the left, mRFP in the middle, and their overlays with bright-field signals on the right. ( C ) Scoring of EGFP and mRFP mutation germline transmission rates. EGFP (left panel) and mRFP (right panel) gRNA-induced mutation germline transmission rates are sorted from lowest (100% wild type) to highest (100% mutant). Each bar represents G1 scoring of an individual cross from a single injected G0 Vasa-Cas9 fly. Twenty-one G0 crosses (total 465 G1 flies) were scored for EGFP and 48 G0 crosses (total 1418 G1 flies) were scored for mRFP gRNA-induced G1 mutations. Yellow portions indicate the percentage of G1 flies expressing both EGFP and mRFP (wild type), red portions indicate mRFP only expression (EGFP mutant), and green portions indicate EGFP only expression (mRFP mutant). Fifteen out of 21 (71%) G0 EGFP-gRNA vector injected adults and 17 out of 48 (35%) G0 mRFP-gRNA vector injected adults harbor transmittable G1 germline mutations. Two EGFP- and 1 mRFP-gRNA vector injected G0adults produced 100 percent G1 mutant progenies.
Article Snippet:
Techniques: CRISPR, Transgenic Assay, Expressing, Plasmid Preparation, Sequencing, Labeling, Injection, Mutagenesis, Transmission Assay, Produced

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) uMAP projection of cell density by control and BRCA1 +/mut fibroblasts. b) Volcano plot with all differentially expressed genes between control and BRCA1 +/mut fibroblasts. Top 50 BRCA1 +/mut genes were used to define pre-CAF signature. Top 50 control genes define our control Fibroblast Signature. c) Gene signature scoring of all control and BRCA1 +/mut fibroblasts for cancer associated fibroblast (CAF) and iCAF signatures. P-value determined by welch two sample t-test. d) Overall survival Kaplan-Meier analysis of n=1764 patients with a high (n=970) and low (n=794) expression of the pre-CAF signature. e) Overall survival Kaplan-Meier analysis of n=1764 patients with a high (n=1197) and low (n=567) expression of the control fibroblast signature. f) Boxplot of MMP3 expression in control and BRCA1 +/mut fibroblasts. P values are determined using a Mann-Whitney test. g) In situ immunofluorescence (IF) analysis of lobular and ductal regions of breast epithelium in control and BRCA1 +/mut human tissue sections. Scale bar = 50 µm. h) Dot plot showing percentage of MMP3-positive stromal cells in control (blue) and BRCA1 +/mut (red) samples as quantified from IF stainings. P -values were determined using unpaired t-tests.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Expressing, MANN-WHITNEY, In Situ, Immunofluorescence

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) Pre-CAF gene signature scoring in fibroblasts from individual patients. Libraries with representation of less than 250 fibroblasts were excluded. P -value was determined by a t-test. a) Example immunofluorescence microscopy images from ductal and lobular regions in control tissue samples. Values represent the quantified percentage of stromal cells positive for MMP3. Scale bar = 50 µm. b) Example immunofluorescence microscopy images from ductal and lobular regions in BRCA1 +/mut tissue samples. Values represent the quantified percentage of stromal cells positive for MMP3. Scale bar = 50 µm.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Immunofluorescence, Microscopy

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) Schematic primary human 3D co-culture using FACS-isolated epithelial cells and fibroblasts modulated using lentiviral transduction. b) Representative images depicting green fluorescent protein (GFP) expression in transduced fibroblasts in close proximity with epithelial organoids (arrow). Scale bar = 100 μm. c) Co-cultures (5 days) of 4,000 breast epithelial cells seeded alone (no fibroblasts) or with 1x10 4 control fibroblasts transduced with lentivirus (+GFP) or transduced to express MMP3 and GFP (+MMP3). Western blots show increased expression of MMP3 in cells and cultured supernatant of +MMP3 fibroblasts (left). Representative merged bright field and GFP images of co-cultures (scale bar= 400 μm) with arrows indicating mammospheres (GFP-negative). Bar charts (right) represent number of mammospheres quantified per well; values expressed as mean ± SD from three separate experiments with three triplicate wells from each. P -values were determined using unpaired t-tests. d) Co-cultures (5 days) of 4,000 breast epithelial cells seeded alone (no fibroblast) or with 1x10 4 BRCA1 +/mut fibroblasts transduced with lentivirus to express CRISPR- Cas9 and MMP3 gRNA (-MMP3) or GFP only (+GFP) vectors. Western blots show decreased expression of MMP3 in cells and medium of MMP3 knock out fibroblast cultures (left). Representative overlay bright field and GFP images of co-cultures (scale bar= 400 μm) with arrows indicating mammospheres (GFP-negative). Bar charts (right) represent number of mammospheres quantified in each well; values expressed as mean ± SD from triplicates of three independent experiments. P - values were determined using unpaired t-tests. e) 1x10 4 FACS-isolated epithelial cells from patient sample “Control 36” were seeded in Matrigel and treated with 0.5 µg/mL or 1 μg/mL recombinant MMP3. Number of mammospheres was quantified after 5 and 10 days. Representative bright field images of mammospheres after 10 days of culture (scale bar= 400 μm). Bar chart values are expressed as mean ± SD from triplicates of three independent experiments. P -values were determined using unpaired t-tests. f) Schematic of mouse model for evaluating effects of stromal fibroblasts (Control 27) on BRCA1-mediated breast tumorigenesis in vivo . g) Images of dissected tumors after six weeks of growth with reported tumor formation efficiencies. Scale bar= 1 cm. P -values were determined using a one- sided Fisher’s Exact test. h) Volumes of dissected tumors. Values are represented as mean ± SD. control n= 4; +GFP n=8, MMP3 n=12. P -values were determined using unpaired t-tests. i) Masses of dissected tumors. Values are represented as mean ± SD. control n= 4; +GFP n=8, +MMP3 n=12. P -values were determined using unpaired t-tests.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Co-Culture Assay, Isolation, Transduction, Expressing, Western Blot, Cell Culture, CRISPR, Knock-Out, Recombinant, In Vivo

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) FACS plots showing gating strategy for isolation of transduced human fibroblasts with GFP fluorescence in forward and side scatter, singlets gate, GFP gate. b) Representative images of primary mammary epithelial cells cultured alone (control) or with primary mammary fibroblasts transduced with lentivirus to express GFP only (+GFP) or GFP and MMP3 (+MMP3) in Matrigel for 5 days. Fibroblasts are distinguished from epithelial spheres (GFP negative) with GFP fluorescence. Scale bar = 400 μm. c) Quantification of spheres. Values are represented as mean ± SD calculated from nine separate wells per group. P-values were determined by unpaired t-tests. d) Mean values of sphere counts pooled from 3 separate experiments from (b) and . Values are represented as mean ± SD. Statistical significance between all groups was determined with a one-way ANOVA test. e) 10x10 4 FACS-isolated epithelial cells from 4 additional patient samples were seeded in seeded in Matrigel and treated with 0.5 µg/mL or 1 μg/mL MMP3 and spheres were counted after 5 and 10 days. Representative bright field images of mammospheres after 10 days of culture (scale bar= 400 μm). Bar chart values are represented as mean ± SD from triplicates from three separate experiments. P - values were determined using unpaired t-tests. f) Bar graph depicting the fold change in sphere number compared to control (dotted red line) after 10 days of culture with human recombinant MMP3. Bar graph values are expressed as mean ± SD from 15 independent experiments total (5 different patient samples with 3 separate experiments each). g) Primary human breast fibroblasts FACS-isolated from patient sample “Control 27” were transduced to express mouse MMP3 (mMMP3) or GFP only. qPCR analysis was performed on the transduced fibroblasts in two separate trials with three replicates per group. Amplification plot is shown with the difference in the normalized reporter value of the experimental reaction minus the normalized reporter value generated by the instrument (ΔRn) on the y-axis and the cycle number on the x-axis.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Isolation, Fluorescence, Cell Culture, Transduction, Recombinant, Amplification, Generated

Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
Figure Lengend Snippet: a) Unbiased clustering using uMAP projection of all patient epithelial cells. Cells are labeled by mammary epithelial cell state classification as indicated. b) uMAP feature plots displaying single cell energy in facetted plots for control (upper plot) and BRCA1 +/mut cells (lower plot). c) scEnergy distributions across basal, luminal 1, and luminal2 cell types from control and BRCA1 +/mut samples. Mean scEnergy values from individual patients are plotted. P-value determined by welch two sample t-test. d) Volcano plot displaying genes differentially expressed between control and BRCA1 +/mut luminal 1 epithelial cells. e) Gene signature scoring of luminal 1 cells from control and BRCA1 +/mut epithelial cells for basal, myoepithelial, luminal 2-AREG, and luminal 2-MUCL1 marker gene signatures. P-value determined by Welch two sample t-test. f) In situ immunofluroescence analysis of KRT14/KRT19-double positive cells of lobular and ductal regions in control and BRCA1 +/mut tissues with representative images shown. Bar chart (bottom left) indicates the percentage of K14/K19-double positive cells per field of view in control (n=6) and BRCA1 +/mut (n=6). Values are expressed as mean ± SD quantified from at least 5 random fields per patient sample. P -value was determined using an unpaired t-test. g) Single cell Western blot (scWB)-based quantification of the percentage of KRT23+ in FACS-isolated luminal epithelial cells from n=3 control, and n=3 BRCA1 +/mut . Images are representative regions of scWB chips post electrophoresis and antibody probing. Bar chart values are represented as mean ± SD from at least 1000 cells/individual; n=3 control, and n=3 BRCA1 +/mut . h) Schematic workflow for analyzing co-transplant tumors from in vivo study described in -i for KRT8 and KRT5 expression. i) Representative immunofluorescence images of sections from tumors produced from +GFP control or +MMP3 fibroblast transplants. Yellow staining indicates tumor cells double positive for KRT8 and KRT5. Scale bar= 50 µm. j) Bar chart is a quantification of the percent of KRT5 cells positive from KRT8. Values are represented as mean ± SD from counts from three different tumors per group with at least five random fields per tumor tissue. P -value was determined by an unpaired t-test.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Labeling, Marker, In Situ, Western Blot, Isolation, Electrophoresis, In Vivo, Expressing, Immunofluorescence, Produced, Staining
Journal: bioRxiv
Article Title: Pre-neoplastic stromal cells drive BRCA1-mediated breast tumorigenesis
doi: 10.1101/2021.10.20.465221
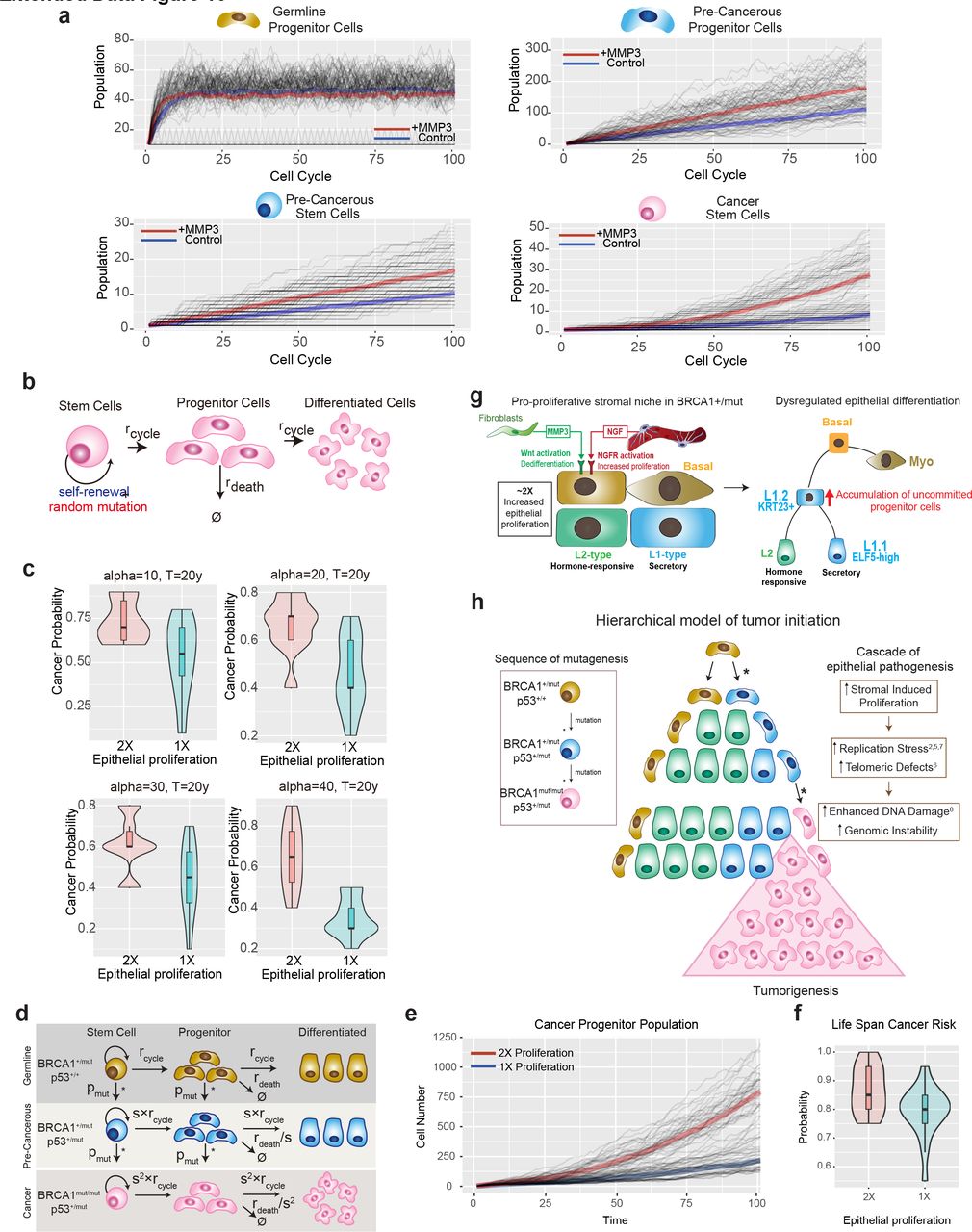
Figure Lengend Snippet: a) The comparison of simulated cell population dynamics in 2 fold increased proliferation and control groups for germline progenitor cells (top left), pre-cancerous progenitor cells (top right), pre-cancerous stem cells (bottom left), and cancer stem cells (bottom right). Thick lines: The averaged population dynamics of 2 fold increased proliferation (red) and control group (blue). Gray thin lines: The stochastic simulation trajectories (sample n=50 for each group). b) Schematic model of the assumptions and parameters used to simulate acquisition of random mutations and consequential fitness change in BRCA1+/mut cells. rcycle is the base line cell division rate, rdeath is the cell date rate. Parameters are further defined in Supplementary Table 10 . c) The robustness of results with respect to parameter alpha in the double-exponential distribution of fitness change ( Supplementary Table 10 ) in the calculation of risk ratio with or without MMP3. The simulation is performed in the time range of 20 years. d) Schematic model illustrating the assumptions and parameters used to simulate the sequential mutations in oncogenes in BRCA1+/mut cells. rcycle is the base line cell division rate, rdeath is the cell date rate, s = proliferation scale factor, pmut is the probability of acquiring a mutation in a driver oncogene. Parameters are further defined in Supplementary Table 11 . e) Comparison between cancer progenitor population dynamics as predicted by a hierarchical model Thick lines: The averaged population dynamics of proliferation in a population with a 2 fold increase in prolifersation and control group (blue). Gray thin lines: The stochastic simulation trajectories (sample n=50 for each group). f) Comparison of the predicted risk ratio of cancer initiation between 2-fold (red) and 1- fold epithelial proliferation rate (blue) in BRCA1 mutation carriers over human life span. The samples are collected from the simulation of N=40 patients in two groups, with the risk ratio of each patient estimated from n=20 simulations of a random mutation model . Violin plots show the distribution of risk ratios over N=20 patients in each group and boxplots indicate median and 25% and 75% quantiles respectively. Wilcoxon Test: *p=0.011. g) Schematic illustrating the concept of a pro-proliferative stromal niche in BRCA1 +/mut breast breast tissues. BRCA1 +/mut stromal cells overepress pro-proliferative cues including NGF in pericytes and pro-tumorigenic MMP3 in fibroblasts (right side). We propose that these act in concert during the pre-neoplastic phase to promote the expansion of a subset of uncommitted luminal progenitor cells as potential cancer cells of origin (left side). h) Concept illustration of the hierarchical model of cancer initiation. Sequence of mutations are indicated in differently colored cells in box on the left, * represent a mutagenic event. Center schematic summarizes outcome of mathematical modeling results indicating expansion of cancer progenitors and ultimately leading to tumorigenesis. Cascade of epithelial cell-intrinsic events promoting tumorigenesis in BRCA1 +/mut is shown on the right. Due to increased stromal cell-induced proliferation and replication stress, BRCA1 +/mut breast epithelial stem cells accumulate mutations and become genomically instable, which increases the likelihood of driver mutations, and ultimately tumor initiation.
Article Snippet: Lentiviral particles were packaged and purchased from
Techniques: Mutagenesis, Sequencing